from IPython.display import HTML
HTML('''
<style>
.nw-anim-wrap{font-family:system-ui,sans-serif;max-width:960px}
.nw-anim-wrap table{border-collapse:collapse;margin:10px 0}
.nw-anim-wrap td,.nw-anim-wrap th{width:38px;height:32px;text-align:center;border:1px solid #999;font-size:13px;padding:2px}
.nw-anim-wrap th{background:#e8e8e8;font-weight:600}
.nw-anim-wrap .cell-unfilled{background:#eee;color:#ccc}
.nw-anim-wrap .cell-filled{background:#fff}
.nw-anim-wrap .cell-current{background:#ffd54f;font-weight:700}
.nw-anim-wrap .cell-diag{background:#90caf9}
.nw-anim-wrap .cell-above{background:#a5d6a7}
.nw-anim-wrap .cell-left{background:#ef9a9a}
.nw-anim-wrap .cell-answer{background:#66bb6a;color:#fff;font-weight:700}
.nw-anim-wrap .cell-trace{background:#ce93d8;font-weight:700}
.nw-anim-wrap .cell-trace-cur{background:#ab47bc;color:#fff;font-weight:700}
.nw-anim-wrap .controls{margin:8px 0;display:flex;align-items:center;gap:8px;flex-wrap:wrap}
.nw-anim-wrap button{padding:4px 14px;font-size:14px;cursor:pointer;border:1px solid #888;border-radius:4px;background:#f5f5f5}
.nw-anim-wrap button:hover{background:#e0e0e0}
.nw-anim-wrap .info-panel{background:#f9f9f9;border:1px solid #ddd;border-radius:6px;padding:10px 14px;margin-top:8px;font-size:13px;line-height:1.6;min-height:60px}
.nw-anim-wrap label{font-size:13px}
.nw-anim-wrap input[type=text]{padding:3px 6px;font-size:13px;width:110px;font-family:monospace}
.nw-anim-wrap input[type=range]{width:100px}
.nw-anim-wrap .step-counter{font-size:13px;color:#555;min-width:90px}
.nw-anim-wrap .alignment-box{font-family:monospace;font-size:14px;margin-top:6px;line-height:1.4}
.nw-anim-wrap .legend{font-size:12px;color:#555;margin-bottom:6px}
.nw-anim-wrap .legend span{display:inline-block;width:14px;height:14px;vertical-align:middle;margin:0 3px 0 8px;border:1px solid #999;border-radius:2px}
</style>
<div class="nw-anim-wrap" id="nwAnimRoot">
<div style="margin-bottom:8px">
<label>x: <input type="text" id="nwInpX" value="TACGTCAGC"></label>
<label style="margin-left:8px">y: <input type="text" id="nwInpY" value="TATGTCATGC"></label>
<button id="nwBtnReset" style="margin-left:8px">Reset</button>
</div>
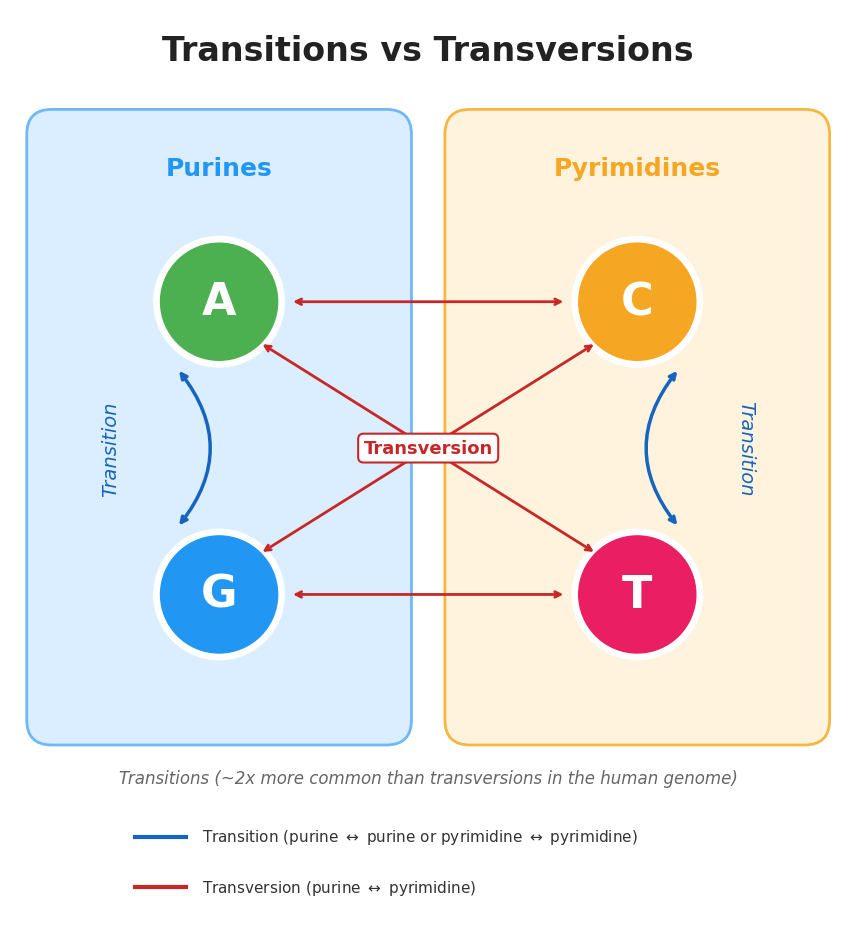
<div class="legend">Penalties: match=0, transition(A/G, C/T)=2, transversion=4, gap=8</div>
<div id="nwGrid"></div>
<div class="controls">
<button id="nwBtnBack">Back</button>
<button id="nwBtnNext">Next</button>
<button id="nwBtnPlay">Play</button>
<span class="step-counter" id="nwStepInfo">Step 0 / 0</span>
<label>Speed: <input type="range" id="nwSpeed" min="1" max="10" value="5"></label>
</div>
<div class="info-panel" id="nwInfo">Press <b>Next</b> or <b>Play</b> to begin.</div>
</div>
<script>
(function(){
function endTag(t){return '<'+'/'+t+'>';}
function esc(s){return s.replace(/&/g,'&').replace(/</g,'<').replace(/>/g,'>');}
var gridDiv=document.getElementById('nwGrid');
var infoDiv=document.getElementById('nwInfo');
var stepInfo=document.getElementById('nwStepInfo');
var inpX=document.getElementById('nwInpX');
var inpY=document.getElementById('nwInpY');
var btnBack=document.getElementById('nwBtnBack');
var btnNext=document.getElementById('nwBtnNext');
var btnPlay=document.getElementById('nwBtnPlay');
var btnReset=document.getElementById('nwBtnReset');
var speedSlider=document.getElementById('nwSpeed');
var steps=[], curStep=-1, D=[], x='', y='', m=0, n=0;
var playing=false, timer=null;
/* Needleman-Wunsch penalty function: match=0, transition=2, transversion=4, gap=8 */
function pen(a,b){
if(a===b) return 0;
if(a==='-'||b==='-') return 8;
var lo=a<b?a:b, hi=a<b?b:a;
if((lo==='A'&&hi==='G')||(lo==='C'&&hi==='T')) return 2;
return 4;
}
function penLabel(a,b){
if(a===b) return 'match';
if(a==='-'||b==='-') return 'gap';
var lo=a<b?a:b, hi=a<b?b:a;
if((lo==='A'&&hi==='G')||(lo==='C'&&hi==='T')) return 'transition';
return 'transversion';
}
function buildSteps(){
x=inpX.value.toUpperCase(); y=inpY.value.toUpperCase();
m=x.length; n=y.length;
D=[];
for(var i=0;i<=m;i++){D[i]=[];for(var j=0;j<=n;j++) D[i][j]=null;}
steps=[];
/* Phase 1: first row — cumulative gap penalties */
D[0][0]=0;
steps.push({type:'init-row',i:0,j:0,val:0,
msg:'Initialize D[0][0] = 0 (both empty strings)'});
for(var j=1;j<=n;j++){
D[0][j]=D[0][j-1]+pen('-',y[j-1]);
steps.push({type:'init-row',i:0,j:j,val:D[0][j],
msg:'Initialize D[0]['+j+'] = D[0]['+(j-1)+'] + gap('+esc(y[j-1])+') = '+D[0][j-1]+' + 8 = '+D[0][j]});
}
/* Phase 2: first column */
for(var i=1;i<=m;i++){
D[i][0]=D[i-1][0]+pen(x[i-1],'-');
steps.push({type:'init-col',i:i,j:0,val:D[i][0],
msg:'Initialize D['+i+'][0] = D['+(i-1)+'][0] + gap('+esc(x[i-1])+') = '+D[i-1][0]+' + 8 = '+D[i][0]});
}
/* Phase 3: fill interior */
for(var i=1;i<=m;i++){
for(var j=1;j<=n;j++){
var pDiag=pen(x[i-1],y[j-1]);
var pAbove=pen(x[i-1],'-');
var pLeft=pen('-',y[j-1]);
var vDiag=D[i-1][j-1]+pDiag;
var vAbove=D[i-1][j]+pAbove;
var vLeft=D[i][j-1]+pLeft;
var best=Math.min(vDiag,vAbove,vLeft);
D[i][j]=best;
var diagDesc=esc(x[i-1])+'/'+esc(y[j-1])+' '+penLabel(x[i-1],y[j-1])+'='+pDiag;
var winners=[];
if(vDiag===best) winners.push('diagonal');
if(vAbove===best) winners.push('above');
if(vLeft===best) winners.push('left');
steps.push({type:'fill',i:i,j:j,val:best,
diag:{i:i-1,j:j-1,val:vDiag},
above:{i:i-1,j:j,val:vAbove},
left:{i:i,j:j-1,val:vLeft},
msg:'D['+i+']['+j+']: diag('+diagDesc+')='+vDiag+
', above(gap)='+vAbove+', left(gap)='+vLeft+
' \u2192 min = <b>'+best+'</b> via '+winners.join(', ')});
}
}
/* Phase 4: answer */
steps.push({type:'answer',i:m,j:n,val:D[m][n],
msg:'Global alignment penalty D['+m+']['+n+'] = <b>'+D[m][n]+'</b>. Next: traceback to recover the alignment.'});
/* Phase 5: traceback */
var tracePath=[];
var ti=m, tj=n;
while(ti>0||tj>0){
tracePath.push({i:ti,j:tj});
if(ti>0&&tj>0){
if(D[ti][tj]===D[ti-1][tj-1]+pen(x[ti-1],y[tj-1])){
ti--;tj--;continue;
}
}
if(ti>0&&D[ti][tj]===D[ti-1][tj]+pen(x[ti-1],'-')){
ti--;continue;
}
tj--;
}
tracePath.push({i:0,j:0});
/* build alignment strings */
var axArr=[], ayArr=[], midArr=[];
for(var k=tracePath.length-1;k>0;k--){
var ci=tracePath[k].i, cj=tracePath[k].j;
var ni=tracePath[k-1].i, nj=tracePath[k-1].j;
if(ni===ci+1&&nj===cj+1){
axArr.push(x[ci]); ayArr.push(y[cj]);
midArr.push(x[ci]===y[cj]?'|':' ');
}else if(ni===ci+1&&nj===cj){
axArr.push(x[ci]); ayArr.push('-'); midArr.push(' ');
}else{
axArr.push('-'); ayArr.push(y[cj]); midArr.push(' ');
}
}
/* traceback steps */
for(var k=0;k<tracePath.length;k++){
var alignHtml='';
if(k>0){
var startIdx=axArr.length-k;
var pAx=axArr.slice(startIdx).join('');
var pMid=midArr.slice(startIdx).join('');
var pAy=ayArr.slice(startIdx).join('');
alignHtml='<div class="alignment-box">'+
'x: '+esc(pAx)+'<br>'+
' '+pMid.replace(/ /g,' ')+'<br>'+
'y: '+esc(pAy)+'</div>';
}
var pi=tracePath[k].i, pj=tracePath[k].j;
var opMsg='';
if(k===0){
opMsg='<b>Traceback</b>: start at D['+pi+']['+pj+'] = '+D[pi][pj];
}else{
var prv=tracePath[k-1];
if(pi===prv.i-1&&pj===prv.j-1){
var lbl=penLabel(x[pi],y[pj]);
if(lbl==='match') opMsg='\u2196 Match: '+esc(x[pi])+'='+esc(y[pj]);
else opMsg='\u2196 '+lbl.charAt(0).toUpperCase()+lbl.slice(1)+': '+esc(x[pi])+'\u2192'+esc(y[pj])+' (cost '+pen(x[pi],y[pj])+')';
}else if(pi===prv.i-1&&pj===prv.j){
opMsg='\u2191 Gap in y: delete '+esc(x[pi])+' (cost 8)';
}else{
opMsg='\u2190 Gap in x: insert '+esc(y[pj])+' (cost 8)';
}
opMsg='<b>Traceback</b> at D['+pi+']['+pj+']: '+opMsg;
}
steps.push({type:'trace',idx:k,path:tracePath,
msg:opMsg+alignHtml});
}
/* final summary */
steps.push({type:'trace-done',path:tracePath,
msg:'<b>Traceback complete!</b> Alignment penalty = <b>'+D[m][n]+'</b>'+
'<div class="alignment-box">'+
'x: '+esc(axArr.join(''))+'<br>'+
' '+midArr.join('').replace(/ /g,' ')+'<br>'+
'y: '+esc(ayArr.join(''))+'</div>'});
}
function render(){
var filled={};
var highlight={};
for(var s=0;s<=curStep&&s<steps.length;s++){
var st=steps[s];
if(st.type==='fill'||st.type==='init-row'||st.type==='init-col'||st.type==='answer'){
filled[st.i+','+st.j]=st.val;
}
}
if(curStep>=0&&steps[curStep].type&&(steps[curStep].type==='trace'||steps[curStep].type==='trace-done')){
for(var i=0;i<=m;i++) for(var j=0;j<=n;j++) filled[i+','+j]=D[i][j];
}
if(curStep>=0&&curStep<steps.length){
var st=steps[curStep];
if(st.type==='fill'){
highlight[st.diag.i+','+st.diag.j]='cell-diag';
highlight[st.above.i+','+st.above.j]='cell-above';
highlight[st.left.i+','+st.left.j]='cell-left';
highlight[st.i+','+st.j]='cell-current';
}else if(st.type==='answer'){
highlight[st.i+','+st.j]='cell-answer';
}else if(st.type==='trace'){
var path=st.path;
for(var k=0;k<=st.idx;k++){
var pk=path[k].i+','+path[k].j;
highlight[pk]=(k===st.idx)?'cell-trace-cur':'cell-trace';
}
}else if(st.type==='trace-done'){
var path=st.path;
for(var k=0;k<path.length;k++){
highlight[path[k].i+','+path[k].j]='cell-trace';
}
highlight[path[0].i+','+path[0].j]='cell-answer';
highlight[path[path.length-1].i+','+path[path.length-1].j]='cell-answer';
}else{
highlight[st.i+','+st.j]='cell-current';
}
}
var h='<table><tr><th>'+endTag('th')+'<th>-'+endTag('th');
for(var j=0;j<n;j++) h+='<th>'+esc(y[j])+endTag('th');
h+=endTag('tr');
for(var i=0;i<=m;i++){
h+='<tr><th>'+(i===0?'-':esc(x[i-1]))+endTag('th');
for(var j=0;j<=n;j++){
var key=i+','+j;
var cls='cell-unfilled';
if(key in filled){
cls=highlight[key]||'cell-filled';
}
var val=(key in filled)?filled[key]:'';
h+='<td class="'+cls+'">'+val+endTag('td');
}
h+=endTag('tr');
}
h+=endTag('table');
gridDiv.innerHTML=h;
if(curStep<0){
infoDiv.innerHTML='Press <b>Next</b> or <b>Play</b> to begin.';
stepInfo.textContent='Step 0 / '+steps.length;
}else{
infoDiv.innerHTML=steps[curStep].msg;
stepInfo.textContent='Step '+(curStep+1)+' / '+steps.length;
}
}
function init(){
stopPlay();
curStep=-1;
buildSteps();
render();
}
function stepFwd(){
if(curStep<steps.length-1){curStep++;render();return true;}
return false;
}
function stepBack(){
if(curStep>=0){curStep--;render();}
}
function getDelay(){return 600-(speedSlider.value-1)*55;}
function startPlay(){
playing=true;btnPlay.textContent='Pause';
function tick(){
if(!playing)return;
if(!stepFwd()){stopPlay();return;}
timer=setTimeout(tick,getDelay());
}
tick();
}
function stopPlay(){playing=false;btnPlay.textContent='Play';if(timer){clearTimeout(timer);timer=null;}}
btnNext.addEventListener('click',function(){stopPlay();stepFwd();});
btnBack.addEventListener('click',function(){stopPlay();stepBack();});
btnPlay.addEventListener('click',function(){if(playing){stopPlay();}else{startPlay();}});
btnReset.addEventListener('click',init);
init();
})();
</script>
''')